
Bioremediation
Remediation Research: Approaches Through Phytoremediation and Synthetic Biology
Phytoremedation is the use of plants and plant/microbe combinations to cheaply clean toxic spills through natural processes. We're working on a specific type of plant/microbe combination, the ectomycorrhizal symbiosis. Ectomycorrhizae are formed by the symbiotic relationship between certain groups of plants (for example California natives such as oaks, willows and pines) and fungi that evolved from those that break down wood, which is high in phenolic compounds. Our research is based on utilizing the natural ability of fungi in this symbiosis to oxidize contaminants based on phenol, such as polycyclic aromatic hydrocarbons (PAH) and polychlorinated benzenes (PCB), and also raw diesel. We have been answering 2 questions. A), how can we increase the efficiency of this naturally-occurring phenomenon? B), which microbes can we use that are specifically adapted to and grow well under chemical conditions created in contaminated soils?
A) How can we quickly and easily increase the efficiency of naturally-occurring bioremedation agents? Ectomycorrhizae are produced by fungi that infiltrate plant roots. In this relationship, the plant gets its nitrogen from the fungus which breaks down phenolic-based compounds in the soils, and the fungus gets it's sugars from that are produced by photosynthesis. Neither partner can survive alone. Our rationale was that if the sugars from the plant were reduced by decreasing how much they could make through phtosynthesis, the fungi might increase their production of enzymes to get their sugars from the surrounding soil, which is very high in phenolic-based compounds. Our method of decreasing photosynthetic capacity was partial defoliation of the plant partner, in this case a pine tree. Our data indicate that we could indeed get the fungal partners to up-regulate their enzyme productions, by up to 10-fold in some cases. This increase occurs whether the defoliation is induced purposely (e.g.,1) or naturally through factors such as disease (e.g., 2). Because we are working in a natural ecosystem, not only do these studies provide fungi and fungal enzymes for remediation, but they also provide organisms that can be used in space flight for waste recycling. Further, our data changed the way we look at symbiosis in general, and provided new data for use in climate change models, Thus, this work is of interest to NASA Earth Sciences and Space Life Support and Sustainability, and also to the broader scientific community.

B) Bioprospecting: Locating fungal enzymes specifically adapted to conditions in contaminated soils. We have performed several experiments in order to locate fungi with enzymes specifically adapted to conditions within contaminants of interest to Ames. In one, we applied natural phenol-based substrates to an ectomycorrhizal system (e.g., 3). This provides enzyme sources for an array of PAH/PCB's. Second, we assayed the natural community in acid-thermal soils. The high acid, high sulfate mimic conditions such as those found in diesel spills, and produced by phenolic oxidation (paper in prep). In the third, we applied diesel to a mycorrhizal soil. This work has provided and continues to provide us with a number of fungai that are good candidates for use in phytoremediation strategies. We have found that fungi in 2 genera do particularly well, for example species in the genus Russula (paper in prep, Russula sp pictured here).
Applications At Ames: Our data provide a solution to soil contamination that is both cheap, and aesthetically pleasing, and also serves to restore Base habitat. This can be done through restoration of na�ve plant communities that naturally possess the proper microbial associations (such as oak or scrub oak) or through the utilization of ornamental plants already present on base (such as the pines) to act as the phytoremediation agents, in conjunction with increasing the rate of remediation by up to 10-fold by simply removing half the leaves or needles. Remediation will occur well beyond the range of the plant�s roots as the fungal structures (hyphae) can extend far beyond the roots. In fact, the largest living organism is a fungus, and the hyphae from a single individual can cover over 2000 acres.
Taking results from the above experiments, I have been funded by NSF (Early Concept Grant for Exploratory Research, award # DEB 1046406 to Cullings awarded 9/2010) to purify enzymes from our candidate fungi for use in remediation, and to continue field studies. Enzymes have several advantages over traditional methods of remediation, including production of less toxic by-products, greater potential to enhance bio-availability of breakdown products for further remediation, and the feasibility of enzyme production at industrial scales.
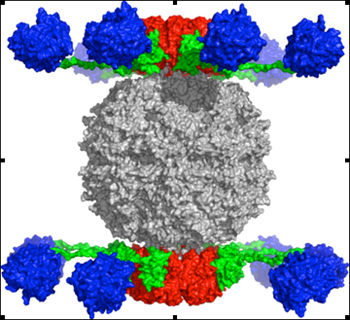
Using enzymes from fungi identified by our above experiments we are providing "parent enzymes" that can be used and/or engineered with greater efficiency under the conditions required for enzyme function in contaminated soil and water. The enzymes we purify in our NSF-funded study will be utilized in partnership with Chad Paavola in the SCB group at Ames, and with the Industrial Sustainable Design group of SCION, a Crown Research Institute of New Zealand to engineer enzymes and enzyme complexes with higher activity. One exciting project is to construct a protein/enzyme combination termed a Rosettasome (pictured) so named because of its Rosetta Stone quality. It is comprised of a protein base upon which different enzymes (in blue) can be placed. The type of enzymes and their physical arrangement can be custom fit to the contaminant of choice. Thus, this is a flexible platform with nearly endless application. It could be applied directly to soils or surfaces, or fixed to a filter through which contaminated groundwater could be pumped. Thus, it will have wide-ranging application to Ames, and to many contaminated systems. We have just submitted a proposal to NSF for further extramural funding for this project.
1) Cullings, K.W., Ishkhanova, G., and J. Henson. 2008. Defoliation effects on enzyme activities of the ectomycorrhizal fungus Suillus granulatus in a Pinus contorta (lodgepole pine) stand in Yellowstone National Park. Oecologia 158(1): 1432-1439
2) Cullings, K and Hanely, J. in press. Dwarf Mistletoe Effects on Soil Basidiomycete Community Structure, Soil Fungal Functional Diversity, and Soil Enzyme Function: Implications for Climate Change. Soil Biology.
3) Cullings, K, Ishkhanova, G., and Ishkhanov, G. in press. Induction of saphophytic behavior in the ectomycorrhizal fungus Suillus granulatus by litter addition in a Pinus controta (Lodgepole pine) stand in Yellowstone. Soil Biology and Biochemistry.